
|
|

|

|
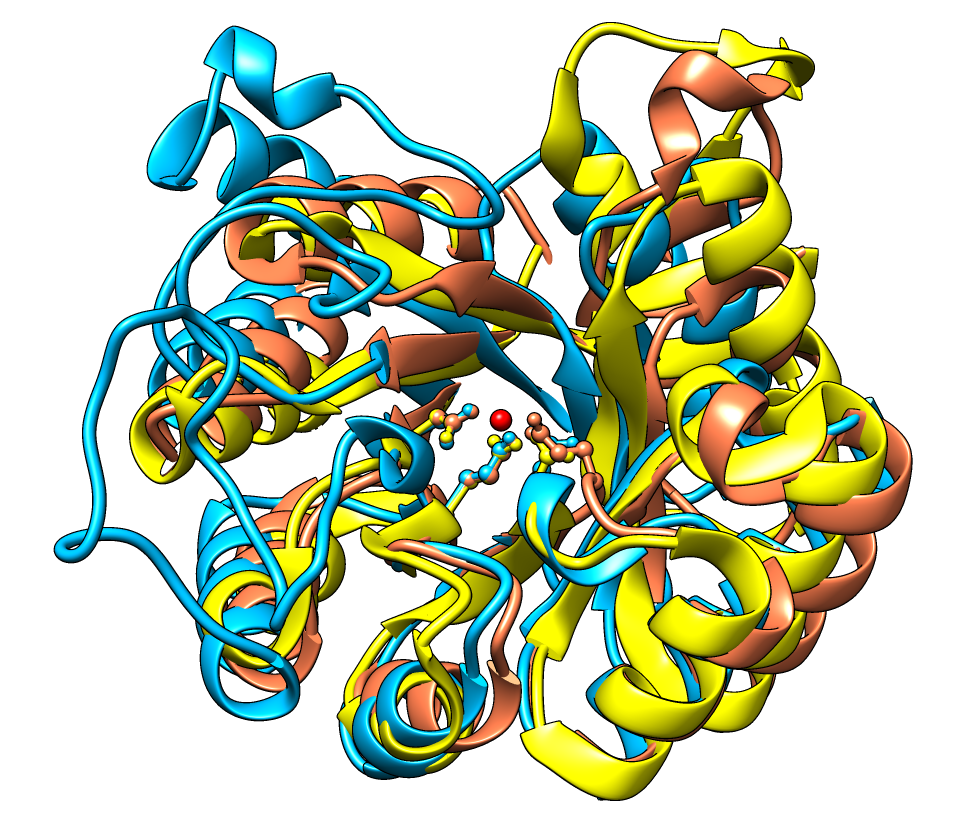
This example from Patricia Babbitt's research group at UCSF shows an alignment of three proteins in the enolase superfamily. The Babbitt group is characterizing superfamilies of enzymes whose member proteins perform a broad range of biochemical functions while sharing a common active site architecture.
Three residues in each protein are shown in ball-and-stick in the structures and highlighted with purple in the sequence viewer. These conserved residues bind a divalent metal ion (shown as a red ball), and along with other residues, they provide the machinery for abstracting a proton α to a carboxylic acid. This partial reaction is the common feature of enolase superfamily members.
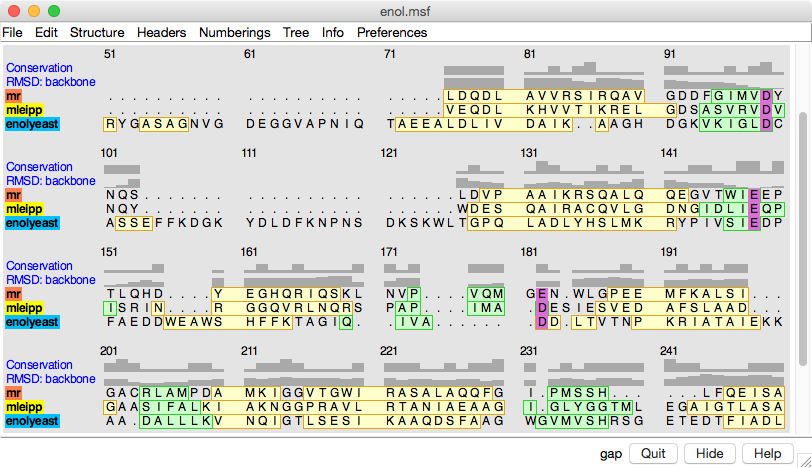
In the sequence alignment, the color swatches behind the sequence names match the corresponding structures, and yellow and green boxes delimit α-helices and β-strands, respectively. Histograms above the sequences show sequence conservation by the AL2CO entropy measure and spatial conservation by backbone atom RMSD.
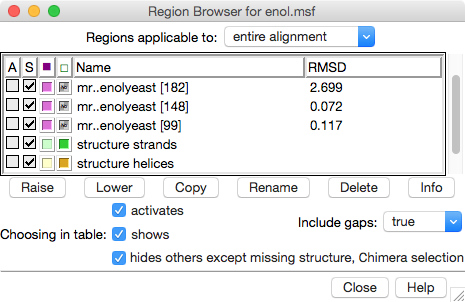
The Region Browser dialog on the right lists the “regions” (colored boxes) in the alignment and allows showing/hiding them.
© 2004, 2015 The Regents of the University of California; all rights reserved.